Mero acaso, fortuita necessidade ou design inteligente???
terça-feira, dezembro 29, 2020
Um megatsunami atingiu a costa israelense há 10.000 anos atrás.
segunda-feira, dezembro 28, 2020
Nanomáquinas propulsoras: evolução convergente de arquelas, flagelos e cílios ou design inteligente?
terça-feira, dezembro 08, 2020
Será que o cérebro humano se parece com o universo? Mero acaso, fortuita necessidade ou design inteligente?
terça-feira, novembro 17, 2020
The Quantitative Comparison Between the Neuronal Network and the Cosmic Web
www.frontiersin.orgF. Vazza1,2,3* and www.frontiersin.orgA. Feletti4,5
1Dipartimento di Fisica e Astronomia, Universitá di Bologna, , Bologna, Italy
2Hamburger Sternwarte, Hamburg, Germany
3Istituto di Radio Astronomia, INAF, Bologna, Italy
4Institute of Neurosurgery, Department of Neurosciences, Biomedicine, and Movement Sciences, University of Verona, Verona, Italy
5Azienda Ospedaliera‐Universitaria di Modena, Modena, Italy
Mais uma hipótese sobre a origem da vida: modelos de precursores potenciais de células resistem às condições simuladas da Terra primitiva
quinta-feira, outubro 29, 2020
Editores de revistas científicas não devem contribuir para politizar a ciência!
segunda-feira, outubro 26, 2020
Science journal editors shouldn’t contribute to politicizing science
By GENEVIEVE P. KANTER OCTOBER 23, 2020
When the editors of some of the world’s leading science journals agree on something, it is generally safe to assume that they are correct. So when prominent journals like Science, Nature, and the New England Journal of Medicine recently published editorials excoriating President Trump’s deadly bungling of the pandemic response and suppression of scientific activity, the editors accurately spotlighted the troubling deficiencies of the current administration.
But in advocating against or endorsing a presidential candidate, these editors made a grave error. In taking this extraordinary step, they made themselves vulnerable to charges of bias, overstepped their roles as science editors, and succumbed to the politicization of science that they and many other scientists find so alarming.
At first glance, these appear to be similar to run-of-the-mill newspaper endorsements. This analogy, however, is not quite right. At a newspaper, there is a wall between the news operation and the editorial office. It exists to prevent biases of the editorial staff from influencing news reporting. No such wall exists for science journals. The editors who write the editorials are the same ones who evaluate manuscripts and make the final decisions on whether to publish them.
There’s another problem: This political advocacy unnecessarily invites allusions to cronyism, echoing a less savory time when wealthy newspaper owners used their editorial pages to extol the merits of their political chums. Indeed, because of fears surrounding the appearance of undue influence and bias, many newspapers in recent years have abandoned political endorsements.
Read more here: STAT News
Transporte de membranas - elevadores condutores para o interior e exterior das células: mero acaso, fortuita necessidade ou design inteligente?
quinta-feira, outubro 15, 2020
Se o Big Bang não foi o começo, foi o que?
terça-feira, setembro 22, 2020
Megalodon, um antigo tubarão extinto de grande dimensão corporal.
segunda-feira, setembro 07, 2020
A estátua de Darwin vai ser demolida por causa do seu livro racista The Descent of Man???
Ajuste fino de máquinas e sistemas moleculares: mero acaso, fortuita necessidade ou design inteligente?
sexta-feira, setembro 04, 2020
Volume 501, 21 September 2020, 110352
Using statistical methods to model the fine-tuning of molecular machines and systems
Steinar Thorvaldsen a Ola Hössjer b
https://doi.org/10.1016/j.jtbi.2020.110352
Under a Creative Commons license Open Access
• Statistical methods are appropriate for modelling fine-tuning.
Abstract
Se não há ancestral comum e nem seleção natural, por que ainda chamamos de evolução?
quinta-feira, setembro 03, 2020
Site sobre bioluminescência
quarta-feira, setembro 02, 2020
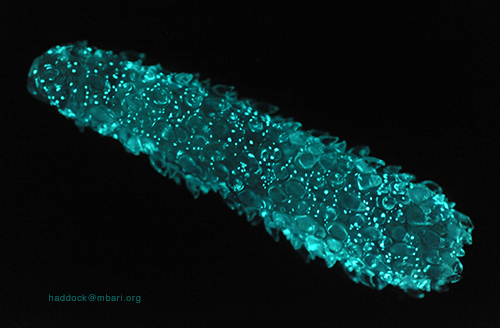
Pyrosomes, colonial salps, continue to be one of the most mysterious of bioluminescent organisms. Their glow can last 15 seconds or more, and it can be triggered by light, even cascading from one end of the colony to the other. The chemical origin remains unknown. It was thought to be bacterial, in part because of similar kinetics, but now appears to be intrinsic chemistry that lets this animal emit its impressive glow.
A ontogenia da nadadeira sarcopterígea elucida a origem das mãos com dedos
quarta-feira, agosto 26, 2020
Darwin, o problema da forma biológica permanece sem solução, mano!
segunda-feira, agosto 17, 2020
On the problem of biological form
Marta Linde-Medina
Theory in Biosciences volume 139, pages299–308(2020)
Abstract
Embryonic development, which inspired the first theories of biological form, was eventually excluded from the conceptual framework of the Modern Synthesis as irrelevant. A major question during the last decades has centred on understanding whether new advances in developmental biology are compatible with the standard view or whether they compel a new theory. Here, I argue that the answer to this question depends on which concept of morphogenesis is held. Morphogenesis can be conceived as (1) a chemically driven or (2) a mechanically driven process. According to the first option, genetic regulatory networks drive morphogenesis. According to the second, morphogenesis results from an invariant tendency of embryonic tissues to restore changes in mechanical stress. While chemically driven morphogenesis allows an extension of the standard view, mechanically driven morphogenesis would deeply transform it. Which of these hypotheses has wider explanatory power is unknown. At present, the problem of biological form remains unsolved.
Subscription or payment needed/Requer assinatura ou pagamento: Theory in Biosciences
Darwin, modelar a biologia evolutiva na física não funciona, mano!

Artigo de 2017 esperava sanar as fissuras/encobrir as rachaduras na biologia evolutiva
Microscopia revisitada
Mais uma hipótese sobre a origem da vida: surgimento de atividade catalítica em um autorreplicador
terça-feira, agosto 04, 2020
Abstract
O impacto de Darwin na sociedade em menos de 3 minutos
sexta-feira, julho 31, 2020
Origem da Vida: o dilema da mão de Deus???
quinta-feira, julho 30, 2020
The Paleobiology Database
sábado, julho 18, 2020

O ajuste fino de máquinas e sistemas moleculares: mero acaso, fortuita necessidade ou design inteligente?
terça-feira, junho 23, 2020
Um padrão irregular de ampulheta descreve o ritmo do desenvolvimento fenotípico na evolução dos mamíferos placentários
segunda-feira, junho 15, 2020
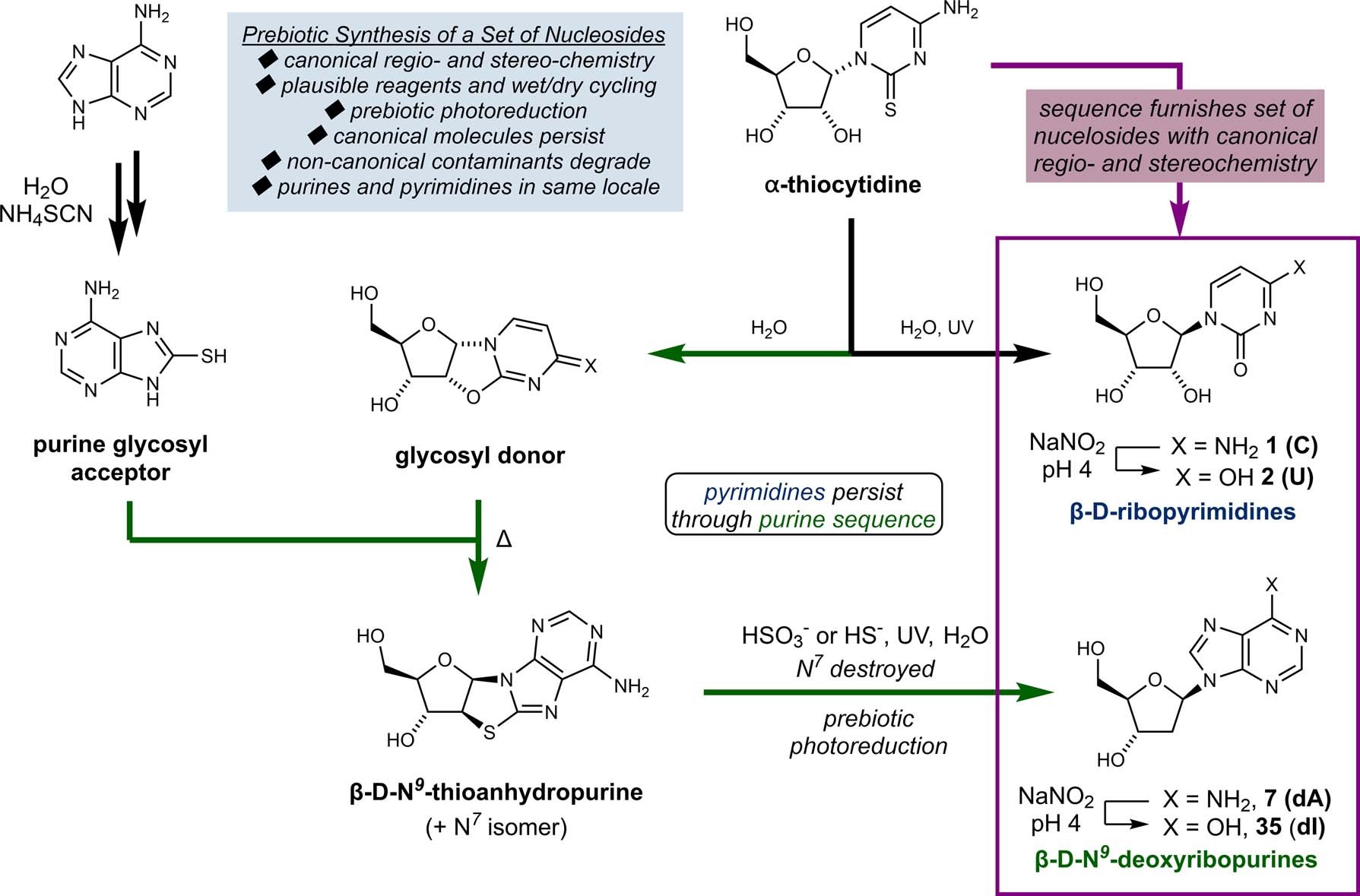
Mais uma hipótese sobre a origem da vida: a partir do DNA e RNA
sexta-feira, junho 12, 2020